18 Dimension Reduction
To represent and visualize high-dimensional data in a meaningful way and help users interpret the data, dimension reduction methods project the data into a lower-dimensional space. Three popular dimension reduction methods are often used: UMAP, PCA, and t-SNE.
Principal Component Analysis (PCA) is a linear dimension reduction method that projects the data onto the directions of the largest variance. It is fast, deterministic, and easier to interpret (PCA loadings are provided), but it is limited by the linear nature of the projection. For a detailed discussion of PCA, please refer to this post.
Uniform Manifold Approximation and Projection (UMAP) and t-Distributed Stochastic Neighbor Embedding (t-SNE) are non-linear dimension reduction methods that can capture more complex patterns in the data. They are more flexible and can handle non-linear relationships between features, but they are more computationally expensive, less interpretable, not deterministic (FLIM Playground uses the random seed 42 to ensure reproducibility), and dependent on the hyperparameters. Here is a post that explains how UMAP and t-SNE work at a high level and the meaning of some of the hyperparameters.
The numerical features are standardized before the dimension reduction.
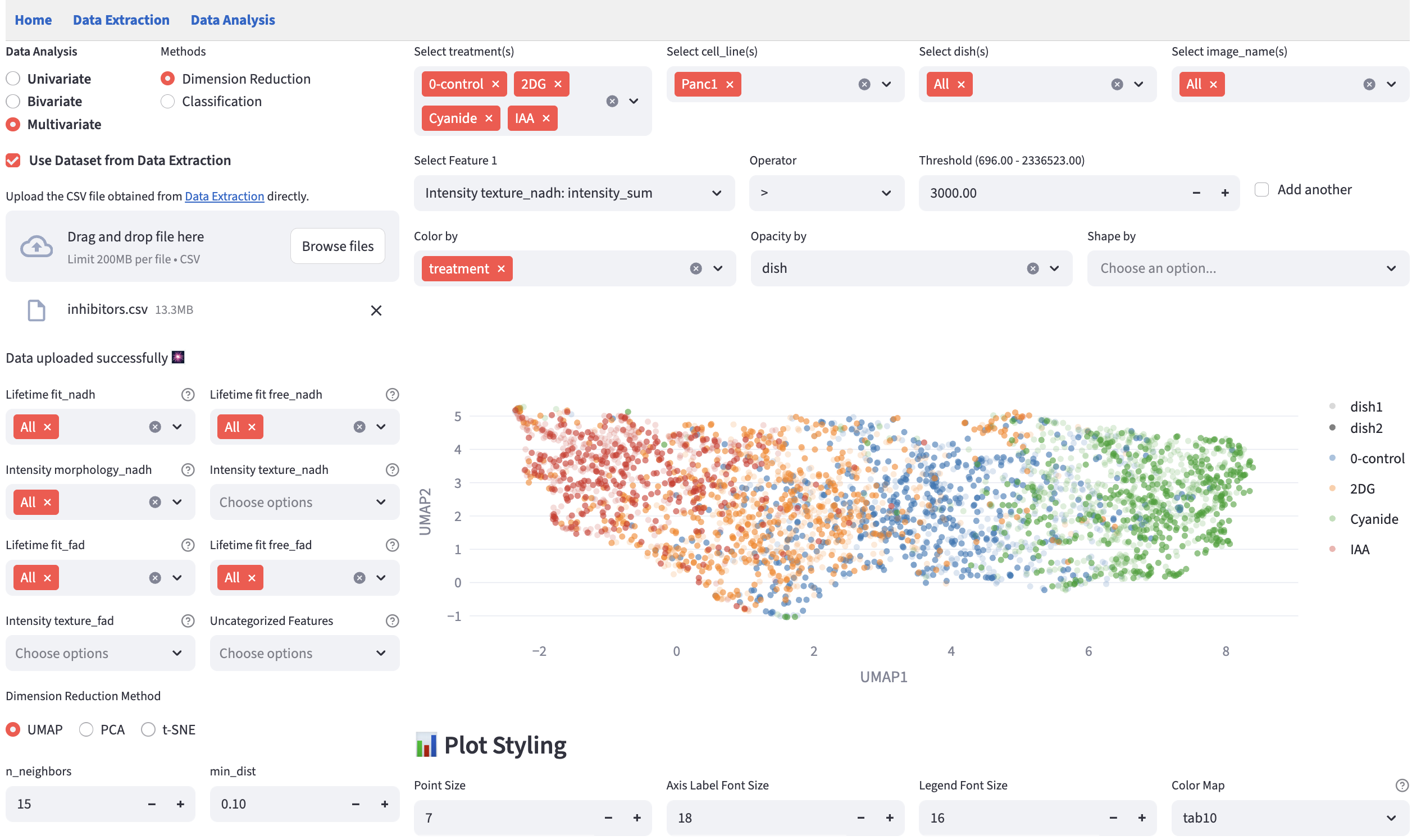
18.2 Hyperparameter Widget
Hyperparameters really matter, so we provide a set of interactive widgets to help users change the hyperparameters and see their effects in real time.
18.2.1 UMAP
FLIM Playground uses the umap-learn implementation of UMAP. It chooses to expose n_neighbors and min_dist for users to tweak. Other hyperparameters are set to default values.
18.2.2 t-SNE
FLIM Playground uses the sklearn.manifold.TSNE implementation of t-SNE. It chooses to expose perplexity and early_exaggeration for users to tweak. Other hyperparameters are set to default values.