3 Configuration
Users can configure the system once, and it will be applied to future data.
Constructing a framework that balances flexibility and usability is about choosing the right abstraction level. FLIM Playground chooses acquisition channel from data levels as the focus and adopts the channel-centric framework. Settings are divided into two categories: cross-channel configurations and per-channel configurations.
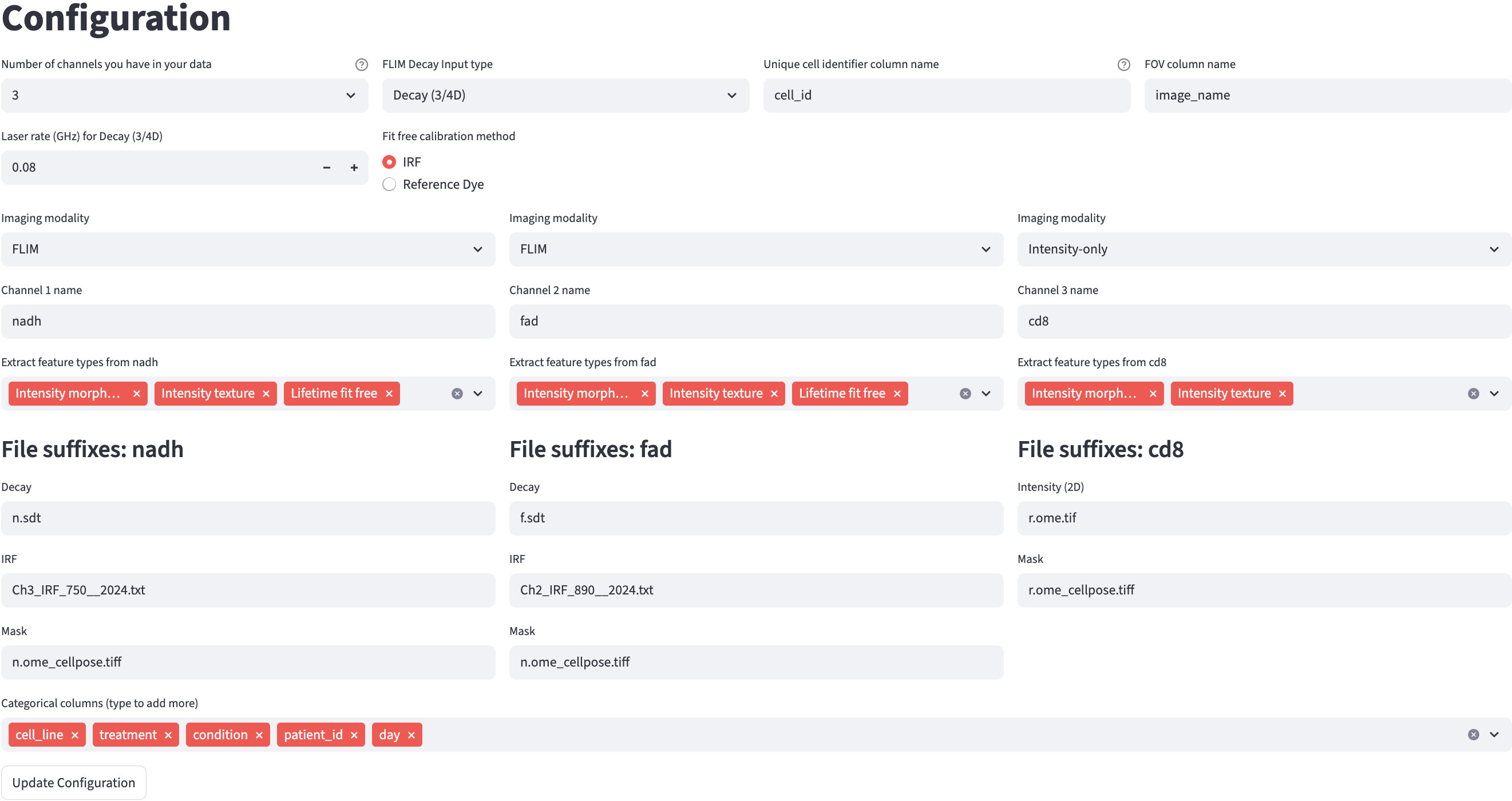
3.1 Cross-Channel Configurations
3.1.1 Categorical Features
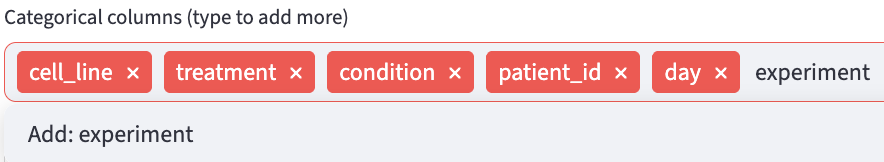
Categorical features are useful to organize the data into groups of interest, and will be used extensively in the Data Analysis section. Users can specify the potential categorical feature names to be extracted in the Categorical Feature Extraction step.
3.1.2 Decay Types
FLIM Playground supports different decay types, including 2D decay, 3D/4D decay, and 3D/4D pixel pre-fit decay, through user specification.
2D Decay
In the system developed by Samimi et al.1, single cells flow through and a decay curve is acquired for each cell by aggregating all the photon arrival time delays deemed to be from the same cell. In exchange for spatial distribution, the acquisition speed is increased enormously. The output of the system is a tabular data sheet with each row representing a cell and each column representing a time bin. FLIM Playground currently supports 2D decay in the CSV format.
3D/4D Decay
In TCSPC-based FLIM, the data is stored in a 3D/4D array and in special format (e.g. .sdt, .ptu). In addition to the spatial dimensions and the time dimension (XYT), the channel dimension may also be present, making it 4-dimensional (CXYT). FLIM Playground currently supports both .sdt and .ptu files up to those 4 dimensions. It reads the duration and the number of time bins from the decay file metadata. Some systems can also record a time stack, which is not supported yet.
3D/4D pixel pre-fit
Since FLIM Playground currently does not support pixel-fitting, users have the option to provide pixel pre-fit outputs from SPCImage in the format of .asc. See here for how FLIM Playground processes SPCImage outputs to obtain cell-level lifetime fitting features.
3.1.3 Identifiers

Cell Identifier
Users can specify the column name for the unique identifier of the cell to be stored in the output dataset. The cell identifier is constructed as {FOV Identifier}_{cell_label}, where the cell_label is the unique integer label in the mask or the row number if the decay type is 2D decay.
FOV Identifier
Users can specify the column name for the field of view identifier. If the decay type is 2D decay, users may want to specify the column name as exp_name, because the FOV is not an image.
3.1.4 Decay Info
Laser frequency
Users can specify the laser frequency in GHz. For example, 0.08 GHz equals 80 MHz. It will be used in the phasor analysis. If you do not intend to use the phasor analysis, you can fill in any number.
2D Decay-specific
Since the 2D decay file does not provide the duration and the number of time bins per laser pulse interval, users need to specify them.

3.1.5 Calibration Method
It supports two calibration methods for phasor analysis:
Since the instrumental response function (IRF) or the fluorescence lifetime standard is measured for each channel, they are set in the per-channel configurations. The fluorescence lifetime standard’s lifetime is channel-agnostic.

3.1.6 Number of Channels
Each FOV is assumed to have the same number of channels. The configuration will dynamically adjust the per-channel configurations based on the specified number of channels. Currently, the maximum number of channels is 4, and it can be extended to house more channels.
3.2 Per-Channel Configurations
3.2.1 Channel Name
Users can customize the channel name based on, for example, the name of the fluorophore.

3.2.2 Imaging Modality
Users can specify the imaging modality for each channel. Currently, two modalities are supported:
FLIM: The signal is time-resolved, i.e., has a time dimension, and is expected to be from one of the decay types.Intensity-only: The signal is a spatial map of intensity values (2 spatial dimensions), i.e., has no time dimension. Currently, the only supported signal type is a 2D intensity image in the format of.tiff/.tif.

This setup allows users to image some channels in FLIM mode, and others in non-FLIM mode. It also allows users to extract and analyze their data if they image in non-FLIM microscopic systems such as epifluorescence, confocal, brightfield, etc. by selecting Intensity-only for all channels.
If the decay type is 2D decay, the imaging modality for all channels is restricted to FLIM.
3.2.3 Feature Extractor
Four types of feature extractors are available if the imaging modality is FLIM:
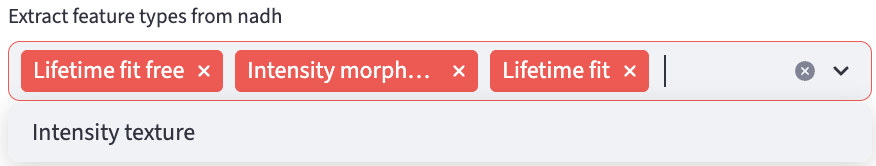
If the decay type is 2D decay, the Intensity Morphology and Intensity Texture feature extractors are not applicable.
The Intensity Morphology and Intensity Texture feature extractors are available if the imaging modality is Intensity-only.
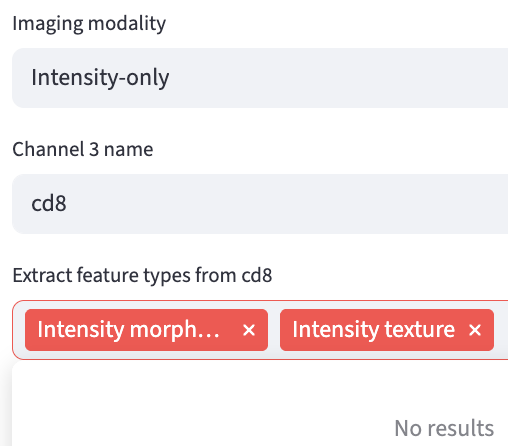
Each channel can be assigned its own set of feature extractors. For example, if users do not intend to do lifetime extraction, they can de-select the Lifetime fit and Lifetime fit free options.
3.2.4 Number of Components
If users select the Lifetime fit under this channel, they can specify the number of components to fit.
3.2.5 Fixed Lifetimes
Users can optionally assign a fixed value (in ns) for specific lifetime components (when \(n>1\), e.g., \(t_1\), \(t_2\)). If a value is provided, it serves as the default fixed lifetime during the interactive fitting phase. If left blank (or set to 0), the lifetime acts as a free parameter to be optimized. This is particularly useful when analyzing fluorophores with prior knowledge of their lifetimes.
3.2.6 File Suffix
The FOV Metadata Organization step looks for associated input files for all channels of a given FOV using file suffixes. Based on the decay type, calibration method, and the selected feature extractors, the system automatically generates the file suffixes for required input files.
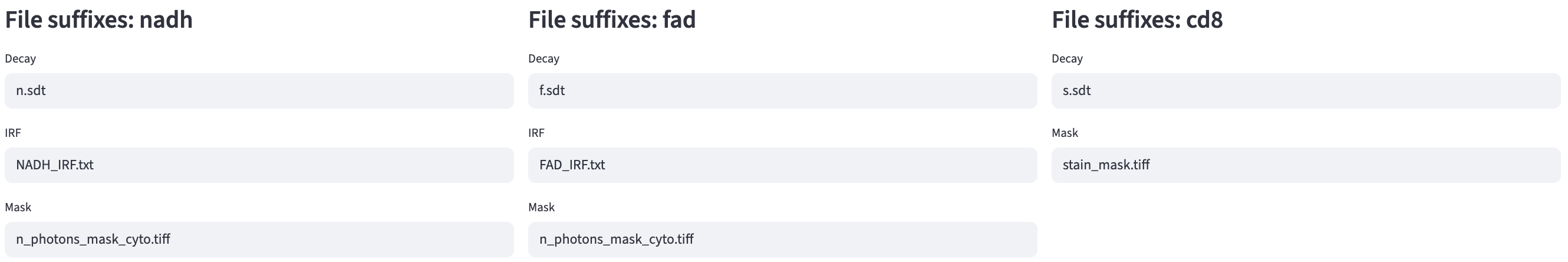
In the example above, channels nadh and fad were asked to provide an IRF file, while channel cd8 did not, because Lifetime fit was selected for the former two channels and not for the latter. The file suffixes of the IRF files were different for each of the two channels, but they shared the same ROI mask file suffix because they used the same ROI mask. The cd8 channel used a different ROI mask than the other two channels, indicated by the mask file suffix.
For detailed information on what file formats are supported for each file type (Mask, IRF, etc.), see input file types.
For detailed information on how the file suffixes are used, and what each file type (e.g. IRF, Mask, etc.) is, see the FOV Metadata Organization page.
3.3 Save
Finally, users can save the configuration for future use by clicking the Update Configuration button.